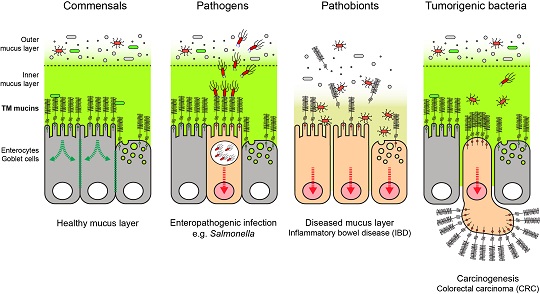
Mucosal microbes in health and disease
Learn about the latest in Mucosal microbes in health and disease. Check out our list of publications.
Stay informed
Get the latest on what is happening in the lab. Check out our news section.